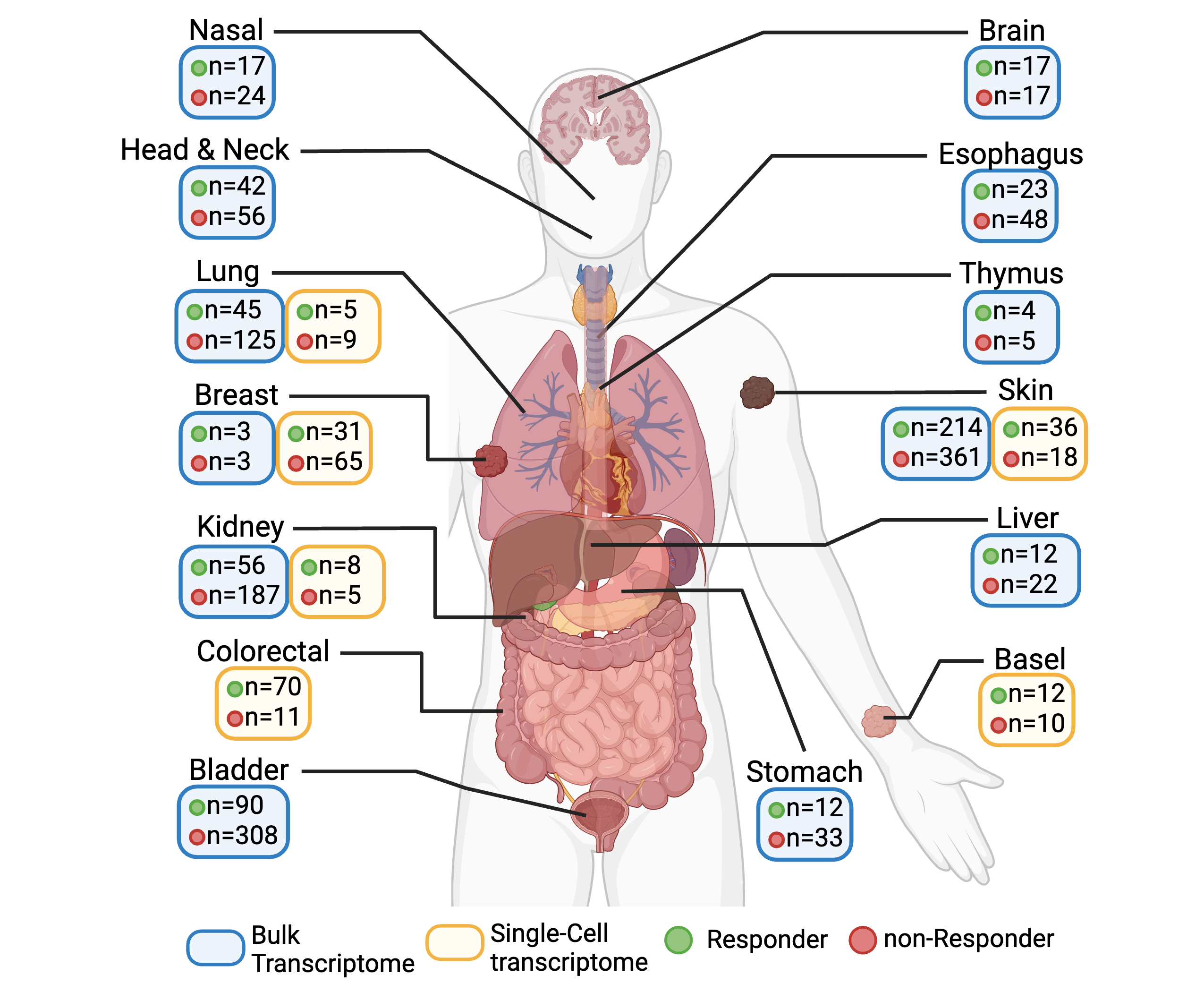
Immuno-Oncology hub (IOhub) is an interactive web server that is built on R Shiny for researchers to investigate immune checkpoint blockade (ICB) treated tumor samples in bulk and single-cell transcriptome data. In IOhub, we have collected 1,806 ICB-treated tumor samples from 36 bulk datasets and 313 samples from 10 single-cell datasets covering 16 cancer types. Please refer to Manual and Resource for detailed information.
This research was supported in part by The National Library of Medicine; National Cancer Institute at the National Institutes of Health grants R00LM013089, R01LM012011 and R01CA254274.
IOhub | © DBMI 2024 | University of Pittsburgh
 Contact: Han Zhang (haz96@pitt.edu), Lujia Chen (luc17@pitt.edu)
Contact: Han Zhang (haz96@pitt.edu), Lujia Chen (luc17@pitt.edu)
Transcriptomic data as well as clinical information were obtained using the public dataset IDs in the below Table. Patients are labeled as 'Response/R' if they showed clinical benefit, complete response (CR), partial response (PR), mixed response (MR) or clonal expansion after ICB treatment; patients are labeled as 'non-Response/NR' if they showed no clinical benefit, progressive disease (PD), stable disease (SD) or no clonal expansion after ICB treatment. Otherwise, samples will be labeled as not evaluated (NE). CR/PD/SD/PD is determined based on RECIST v1.1[1].
Note: The normalized expression datasets are available upon request.
Reference:
[1] Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45, 228-247, doi:10.1016/j.ejca.2008.10.026 (2009).
[2] Hugo, W. et al. Genomic and Transcriptomic Features of Response to Anti-PD-1 Therapy in Metastatic Melanoma. Cell 165, 35-44, doi:10.1016/j.cell.2016.02.065 (2016).
[3] Nathanson, T. et al. Somatic Mutations and Neoepitope Homology in Melanomas Treated with CTLA-4 Blockade. Cancer Immunol Res 5, 84-91, doi:10.1158/2326-6066.CIR-16-0019 (2017).
[4] Riaz, N. et al. Tumor and Microenvironment Evolution during Immunotherapy with Nivolumab. Cell 171, 934-949 e916, doi:10.1016/j.cell.2017.09.028 (2017).
[5] Gide, T. N. et al. Distinct Immune Cell Populations Define Response to Anti-PD-1 Monotherapy and Anti-PD-1/Anti-CTLA-4 Combined Therapy. Cancer Cell 35, 238-255 e236, doi:10.1016/j.ccell.2019.01.003 (2019).
[6] Liu, D. et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat Med 25, 1916-1927, doi:10.1038/s41591-019-0654-5 (2019).
[7] Amato, C. M. et al. Pre-Treatment Mutational and Transcriptomic Landscape of Responding Metastatic Melanoma Patients to Anti-PD1 Immunotherapy. Cancers (Basel) 12, doi:10.3390/cancers12071943 (2020).
[8] Auslander, N. et al. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat Med 24, 1545-1549, doi:10.1038/s41591-018-0157-9 (2018).
[9] Zappasodi, R. et al. CTLA-4 blockade drives loss of T(reg) stability in glycolysis-low tumours. Nature 591, 652-658, doi:10.1038/s41586-021-03326-4 (2021).
[10] Mariathasan, S. et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544-548, doi:10.1038/nature25501 (2018).
[11] Rose, T. L. et al. Fibroblast growth factor receptor 3 alterations and response to immune checkpoint inhibition in metastatic urothelial cancer: a real world experience. Br J Cancer 125, 1251-1260, doi:10.1038/s41416-021-01488-6 (2021).
[12] Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801-806 (2018).
[13] Braun, D. A. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat Med 26, 909-918, doi:10.1038/s41591-020-0839-y (2020).
[14] Shiuan, E. et al. Clinical Features and Multiplatform Molecular Analysis Assist in Understanding Patient Response to Anti-PD-1/PD-L1 in Renal Cell Carcinoma. Cancers (Basel) 13, doi:10.3390/cancers13061475 (2021).
[15] Cho, J. W. et al. Genome-wide identification of differentially methylated promoters and enhancers associated with response to anti-PD-1 therapy in non-small cell lung cancer. Exp Mol Med 52, 1550-1563, doi:10.1038/s12276-020-00493-8 (2020).
[16] Jung, H. et al. DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load. Nat Commun 10, 4278, doi:10.1038/s41467-019-12159-9 (2019).
[17] Obradovic, A. et al. Immunostimulatory Cancer-Associated Fibroblast Subpopulations Can Predict Immunotherapy Response in Head and Neck Cancer. Clin Cancer Res 28, 2094-2109, doi:10.1158/1078-0432.CCR-21-3570 (2022).
[18] Zhao, J. et al. Immune and genomic correlates of response to anti-PD-1 immunotherapy in glioblastoma. Nat Med 25, 462-469, doi:10.1038/s41591-019-0349-y (2019).
[19] Kim, S. T. et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat Med 24, 1449-1458, doi:10.1038/s41591-018-0101-z (2018).
[20] Quintela-Fandino, M. et al. Immuno-priming durvalumab with bevacizumab in HER2-negative advanced breast cancer: a pilot clinical trial. Breast Cancer Res 22, 124, doi:10.1186/s13058-020-01362-y (2020).
[21] van den Ende, T. et al. Neoadjuvant Chemoradiotherapy Combined with Atezolizumab for Resectable Esophageal Adenocarcinoma: A Single-arm Phase II Feasibility Trial (PERFECT). Clin Cancer Res 27, 3351-3359, doi:10.1158/1078-0432.CCR-20-4443 (2021).
[22] He, Y., Ramesh, A., Gusev, Y., Bhuvaneshwar, K. & Giaccone, G. Molecular predictors of response to pembrolizumab in thymic carcinoma. Cell Rep Med 2, 100392, doi:10.1016/j.xcrm.2021.100392 (2021).
[23] Sade-Feldman, M. et al. Defining T Cell States Associated with Response to Checkpoint Immunotherapy in Melanoma. Cell 175, 998-1013 e1020, doi:10.1016/j.cell.2018.10.038 (2018).
[24] Alvarez-Breckenridge, C. et al. Microenvironmental landscape of human melanoma brain metastases in response to immune checkpoint inhibition. Cancer immunology research 10, 996-1012 (2022).
[25] Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell, doi:10.1016/j.ccell.2021.09.010 (2021).
[26] Bassez, A. et al. A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer. Nat Med 27, 820-832, doi:10.1038/s41591-021-01323-8 (2021).
[27] Krishna, C. et al. Single-cell sequencing links multiregional immune landscapes and tissue-resident T cells in ccRCC to tumor topology and therapy efficacy. Cancer Cell 39, 662-677 e666, doi:10.1016/j.ccell.2021.03.007 (2021).
[28] Bi, K. et al. Tumor and immune reprogramming during immunotherapy in advanced renal cell carcinoma. Cancer Cell 39, 649-661 e645, doi:10.1016/j.ccell.2021.02.015 (2021).
[29] Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat Med 25, 1251-1259, doi:10.1038/s41591-019-0522-3 (2019).
[30] Li, J. et al. Remodeling of the immune and stromal cell compartment by PD-1 blockade in mismatch repair-deficient colorectal cancer. Cancer Cell, doi:10.1016/j.ccell.2023.04.011 (2023).
Bulk-RNAseq
For each bulk-RNAseq dataset, gene expressions were normalized in transcript per million (TPM) formats. Samples from CM-009 cohort in Braun’s study were removed as they are duplicated in Miao’s study. In total, 1806 ICB treated tumor samples are included (1496 pre-treatment samples, 310 post-treatment samples; 535 response samples, 1189 non-response samples). We then used non-paranormal normalization (NPN) to remove batch effects and covariates of cancer type. TIDE score[1], IMPRES score[2], Interferon-gamma signature[3], Inflammatory signature[4], GEP inflammation[5], microstallite instability score[6] and tertiary lymphoid structure score[7] were calculated on corrected samples. Meanwhile, Cibersort absolute module[8], xCell[9], EPIC[10] and MCP counter[11] were used to deconvolute the tumor infiltrated lymphocytes in the tumor microenvironment.
scRNAseq
As for scRNAseq datasets, we filtered the cells using the same criteria if 1) the number of genes is less than 250 or higher than 8000; 2) the mitochondrial ratio is greater than 20%; 3) the number of unique molecular identifier (UMI) counts is less than 500. To rule out the bias from manual annotation, we used SingleR[11] to map the cell major type and subtype to Monaco and DICE immune database. In total, we collected 593888 responsive and 280564 non-responsive single cells. The relative proportion of each cell major type and subtype was then summarized for each sample.
Reference:
[1] Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24, 1550-1558, doi:10.1038/s41591-018-0136-1 (2018).
[2] Auslander, N. et al. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat Med 24, 1545-1549, doi:10.1038/s41591-018-0157-9 (2018).
[3] Ayers, M. et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest 127, 2930-2940, doi:10.1172/JCI91190 (2017).
[4] Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, doi:10.1126/science.aaf8399 (2017).
[5] Spranger, S. & Gajewski, T. F. Tumor-intrinsic oncogene pathways mediating immune avoidance. Oncoimmunology 5, e1086862, doi:10.1080/2162402X.2015.1086862 (2016).
[6] Fu, Y. et al. A qualitative transcriptional signature for predicting microsatellite instability status of right-sided Colon Cancer. BMC Genomics 20, 769, doi:10.1186/s12864-019-6129-8 (2019).
[7] Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577, 561-565 (2020).
[8] Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12, 453-457, doi:10.1038/nmeth.3337 (2015).
[9] Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome biology 18, 1-14 (2017).
[10] Racle, J. & Gfeller, D. EPIC: a tool to estimate the proportions of different cell types from bulk gene expression data. Bioinformatics for Cancer Immunotherapy: Methods and Protocols, 233-248 (2020).
[11] Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome biology 17, 1-20 (2016).
02/2025: v0.2 coming soon, with updates in DE and survival analysis.
11/2024: v0.1 released