Overview
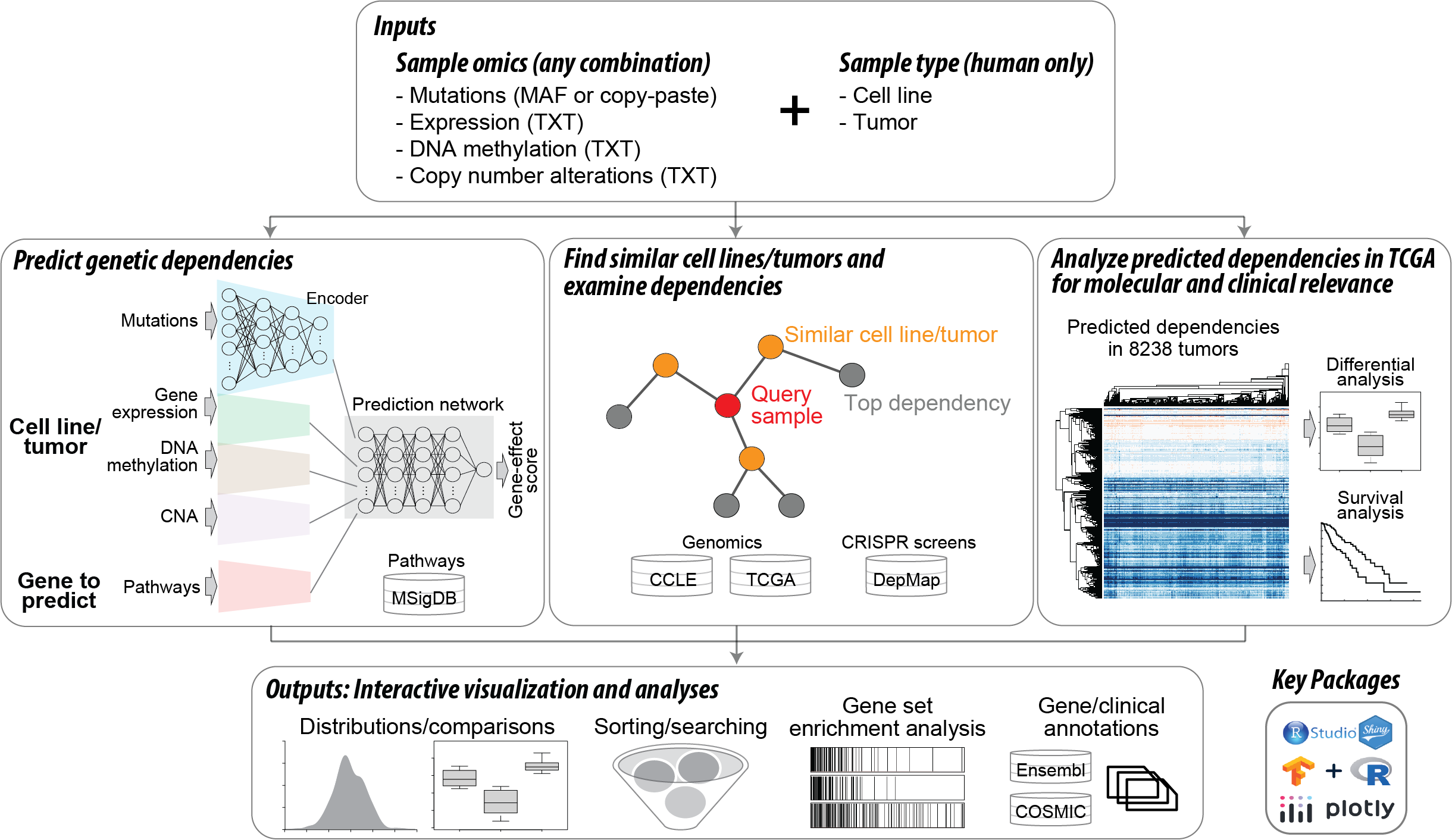
DeepDEP is a deep learning model that predicts gene dependencies using single or multi-omics data of DNA mutation, gene expression, DNA methylation, and copy number alteration (CNA) profiles in a human cancer sample (cell line or tumor) (Chiu et al., Science Advances 2021). The term "genetic dependency" denotes the degree to which the viability of cancer cells is inhibited by knocking out a gene. shinyDeepDEP is an R Shiny app that provides a user-friendly interface to DeepDEP with no requirement of programming skills or computational resources. shinyDeepDEP is designed to predict, prioritize, analyze, and interpret genetic dependencies with it three main analysis modules, Module 1: Predict Genetic Dependencies (core module), Module 2: Find Similar Samples, and Module 3: Analyze TCGA Predictions. Here we provide an overview of our tool. For detailed information regarding the input data format or each module, please visit the corresponding help pages or click on the question mark at each input/output section.
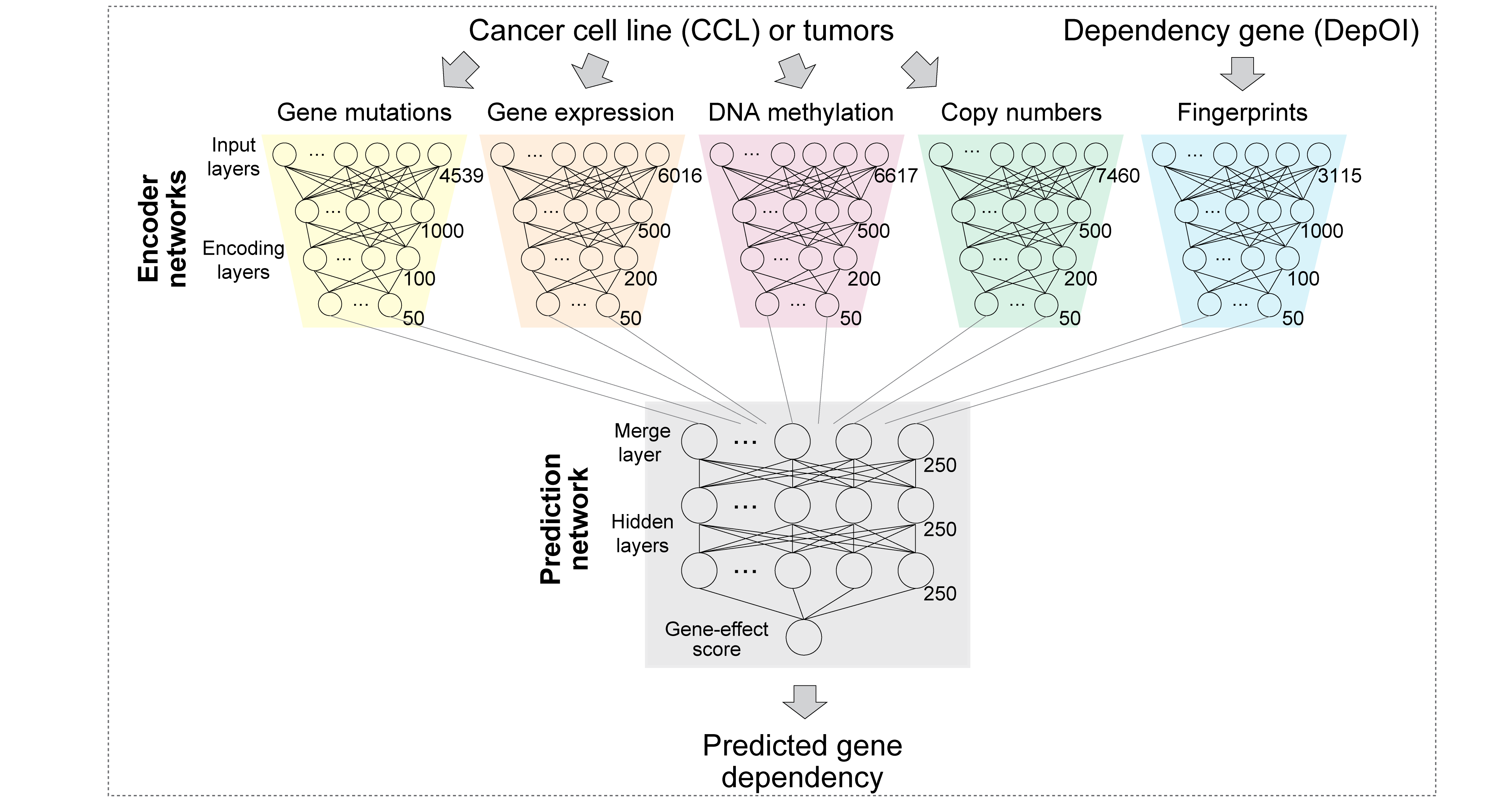
DeepDEP: Deep Learning-Based Prediction of Gene Dependencies of Tumors from Integrated Genomic Profiles
Given omics profiles, DeepDEP predicts gene dependency values for 1,298 pre-selected genes relevant to cancer or genes in a user-selected gene set/pathway. It comprises five component networks:
- A mutation encoder, designed to capture meaningful embeddings of gene mutation data
- An expression encoder
- A DNA methylation encoder
- A copy number alteration (CNA) encoder
- A fingerprints encoder, designed to capture pathway information of a genetic dependency under study
In the publication of DeepDEP, we systematically evaluated the prediction performance using hold-out cancer cell lines and multiple measures, including mean squared error in gene dependency scores and per-cancer cell line correlation coefficients between actual and predicted data. DeepDEP has achieved marked improvements over a diverse array of conventional methods, such as linear regression, support vector machines, and alternative deep learning models trained either with cell lines alone, without transferring features learned from tumors, or using principal components to replace encoder outputs. Furthermore, we validated the predictions with real-world clinical data from corresponding patients. For more details on the model design, validation, and applications, please refer to the publication of DeepDEP (Chiu et al., Science Advances 2021). The Python implementation of DeepDEP is available in our Code Ocean repository.
Along with the full DeepDEP model utilizing all four omics, we have also developed models using only mutation and expression data, as well as a model for each single omics data type. shinyDeepDEP automatically selects the best model based on the data users upload, in the order of full four omics (full DeepDEP), mutations + expression, methylation alone, expression alone, mutations alone, and CNA alone.
shinyDeepDEP: Implementation and Enhancement of DeepDEP by a User-friendly R Shiny Framework
"Module 1: Predict Dependencies (core module)" predicts, prioritizes, and interprets the gene dependencies of the query cancer sample.
This module employs DeepDEP to predict genetic dependencies across the default list of 1,298 cancer-relevant genes or genes involved in a specific gene set/pathway of interest. The R Shiny app offers an intuitive user interface that enables interactive visualization, searching, filtering, and functional interpretation of all prediction results, along with detailed annotations for individual genes.
"Module 2: Find Samples" identifies cell lines/tumors in our curated datasets with similar omics profiles to those of the query sample and examines their genetic dependencies.
This module finds cancer cell lines or tumors in our curated datasets with similar mutation and/or gene expression profiles to those of a query sample and examines the top similar samples' real (for cell lines with CRISPR screening results from DepMap) and predicted genetic dependencies (for tumors with predictions by the full DeepDEP model).
"Module 3: Analyze TCGA" provides access to the pre-generated gene dependencies of TCGA tumors and examines the molecular and clinical relevance.
This module allows users to effortlessly explore pre-calculated gene dependency scores for over 8,000 TCGA tumors as predicted by the full DeepDEP model. It facilitates the analysis of these gene dependencies in conjunction with genetic alterations, such as gene mutations and expression dysregulation, and patient survival, to uncover molecular and clinical insights.